Bivariate analysis#
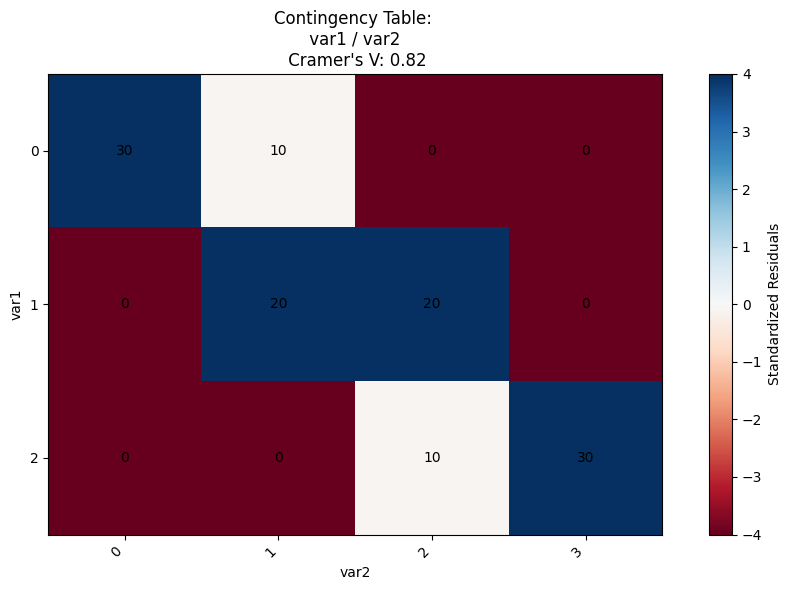
Simple and efficient contingency table with Cramer’s V & standardized residuals#
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import polars as pl
import scipy.stats as stats
import statsmodels.api as sm
df = pd.DataFrame(
{
"var1": [str(i // 40) for i in range(120)],
"var2": [str(i // 30) for i in range(120)],
}
)
def select_dtype(df: pl.DataFrame, dtype: pl.DataType) -> pl.DataFrame:
"""
Select columns with a specific data type using Polars.
"""
# Select "i8" dtype columns
col_dtypes = {df.columns[i]: dtype for i, dtype in enumerate(df.dtypes)}
return df.select(
[pl.col(col) for col, dtype in col_dtypes.items() if dtype == pl.Int8]
)
def get_cramer_v(dataframe: pd.DataFrame, col1: str, col2: str) -> float:
crosstab = pd.crosstab(
dataframe[col1].astype(str),
dataframe[col2].astype(str),
)
crosstab_np = crosstab.to_numpy()
X2 = stats.chi2_contingency(crosstab_np, correction=False)[0]
N = crosstab_np.sum()
min_dim = min(crosstab_np.shape) - 1
cramer_V = np.sqrt(X2 / (N * min_dim))
return float(cramer_V)
def plot_contingency_table(dataframe: pd.DataFrame, col1: str, col2: str):
"""
Plot a contingency table with the given columns.
"""
crosstab = pd.crosstab(
dataframe[col1].astype(str),
dataframe[col2].astype(str),
)
cramer_v = get_cramer_v(dataframe, col1, col2)
contingency_table = sm.stats.Table(crosstab)
standard_resitudals = contingency_table.standardized_resids
plt.figure(figsize=(10, 6))
im = plt.imshow(standard_resitudals, cmap="RdBu", vmin=-4, vmax=4)
for i in range(len(standard_resitudals.index)):
for j in range(len(standard_resitudals.columns)):
_ = plt.text(
j, i, f"{crosstab.to_numpy()[i, j]:.0f}", ha="center", va="center"
)
plt.xticks(
range(len(standard_resitudals.columns)),
standard_resitudals.columns,
rotation=45,
ha="right",
)
plt.yticks(range(len(standard_resitudals.index)), standard_resitudals.index)
plt.title(f"Contingency Table: \n {col1} / {col2} \n Cramer's V: {cramer_v:.2f}")
plt.xlabel(col2)
plt.ylabel(col1)
plt.colorbar(im, label="Standardized Residuals")
plt.tight_layout()
plt.show()
plot_contingency_table(
df,
col1="var1",
col2="var2",
)
Xi correlation#
This type of correlation can be used to measure non-linear and non-monotonic relationships.
import numpy as np
from scipy.stats import rankdata, norm
from typing import Union
def xicor(
x: np.ndarray,
y: np.ndarray,
ties: Union[str, bool] = "auto",
) -> tuple[float, float]:
"""Calculate the correlation coefficient and p-value using the xicor method.
See: https://arxiv.org/abs/1909.10140
Args:
x: First array of numeric values
y: Second array of numeric values
ties: Strategy for handling ties. "auto" detects ties automatically,
or boolean to explicitly enable/disable tie handling
Returns:
Tuple containing:
- correlation statistic (float between -1 and 1)
- p-value for the correlation test
Raises:
ValueError: If input arrays are empty or contain non-numeric values
IndexError: If input arrays have different lengths
TypeError: If ties parameter is invalid type
"""
# Validate and prepare input arrays
if not len(x) or not len(y):
raise ValueError("Input arrays cannot be empty")
x = np.asarray(x, dtype=np.float64).flatten()
y = np.asarray(y, dtype=np.float64).flatten()
n = len(y)
if len(x) != n:
raise IndexError(f"x, y length mismatch: {len(x)}, {len(y)}")
# Handle ties parameter
if ties == "auto":
ties = len(np.unique(y)) < n
elif not isinstance(ties, bool):
raise ValueError(
f'Expected ties to be "auto" or boolean, '
f"got {ties} ({type(ties)}) instead"
)
# Sort y values based on x order and calculate ranks
y_sorted = y[np.argsort(x)]
ranks = rankdata(y_sorted, method="ordinal")
nominator = np.sum(np.abs(np.diff(ranks)))
# Calculate denominator based on ties handling
if ties:
rank_max = rankdata(y_sorted, method="max")
denominator = 2 * np.sum(rank_max * (n - rank_max))
nominator *= n
else:
denominator = np.power(n, 2) - 1
nominator *= 3
# Calculate final statistics
statistic = 1 - nominator / denominator # upper bound is (n - 2) / (n + 1)
p_value = norm.sf(statistic, scale=2 / 5 / np.sqrt(n))
return statistic, p_value
import plotly.express as px
from scipy.stats import pearsonr
nb_points = 100
x = np.linspace(-10, 10, nb_points)
funcs = {
"X, high dispersion": x + np.random.normal(0, 20, nb_points),
"X, low dispersion": x + np.random.normal(0, 5, nb_points),
"X²": np.pow(x, 2) + np.random.normal(0, 10, nb_points),
"X4": np.pow(x / 4, 4)
- 0.075 * np.pow(2 * x, 2)
+ np.random.normal(0, 1.5, nb_points),
"sin": np.sin(x) + np.random.normal(0, 0.5, nb_points),
}
for func, y in funcs.items():
pearson_correlation, p_value = pearsonr(x, y)
xi_correlation, p_value = xicor(x, y)
(
px.scatter(
x=x,
y=y,
title=f"<b>{func}<br></b>Pearson correlation: {pearson_correlation:.2f}<br>Xi correlation: {xi_correlation:.2f}",
)
.update_layout(title_x=0.5)
.show("notebook")
)
